Rigidity-Aware Geometric Pretraining for
Protein Design and Conformational Ensembles
ICLR 2026
- Zhanghan Ni1 *
- Yanjing Li2 *
- Zeju Qiu3 *
- Bernhard Schölkopf3
- Hongyu Guo4,5
- Weiyang Liu3,6
- Shengchao Liu6
- 1University of Illinois Urbana-Champaign
- 2University of Washington
- 3MPI for Intelligent Systems, Tübingen
- 4National Research Council Canada
- 5University of Ottawa
- 6The Chinese University of Hong Kong
- *Equal contribution
1. RigidSSL use inertial frames to establish a canonical reference system for protein structures.
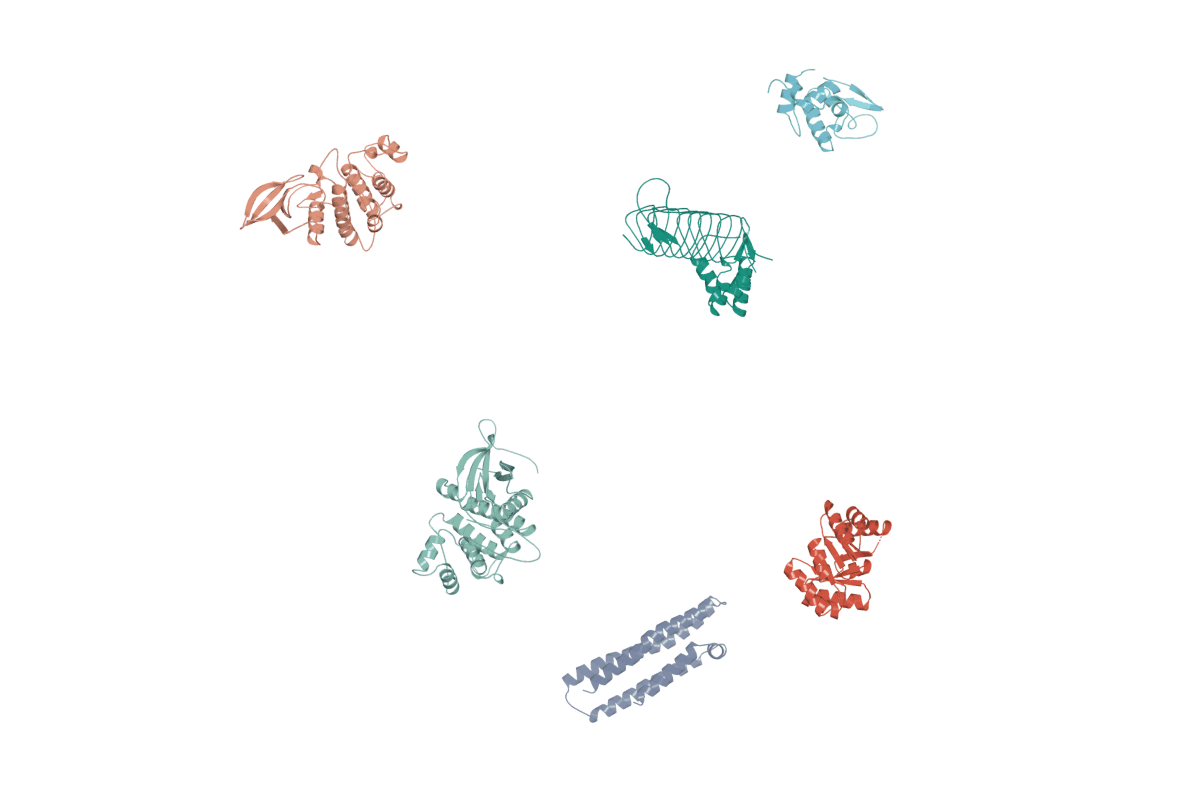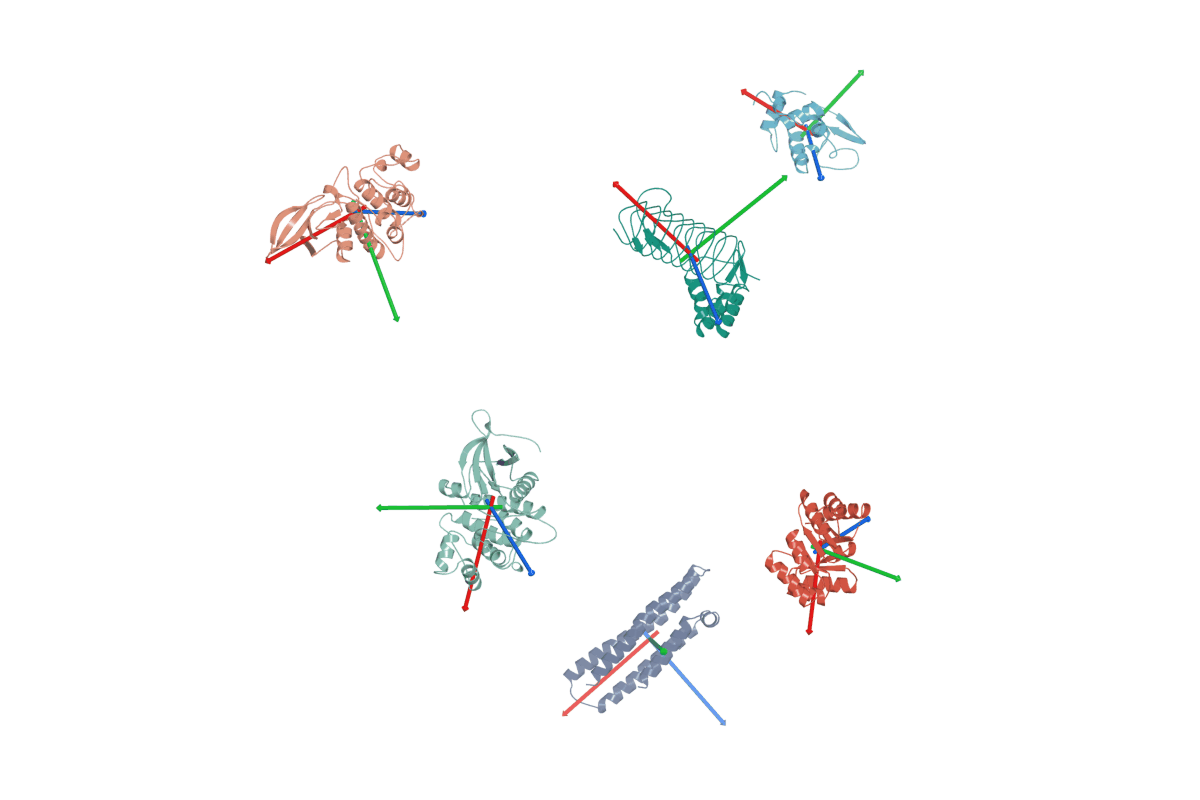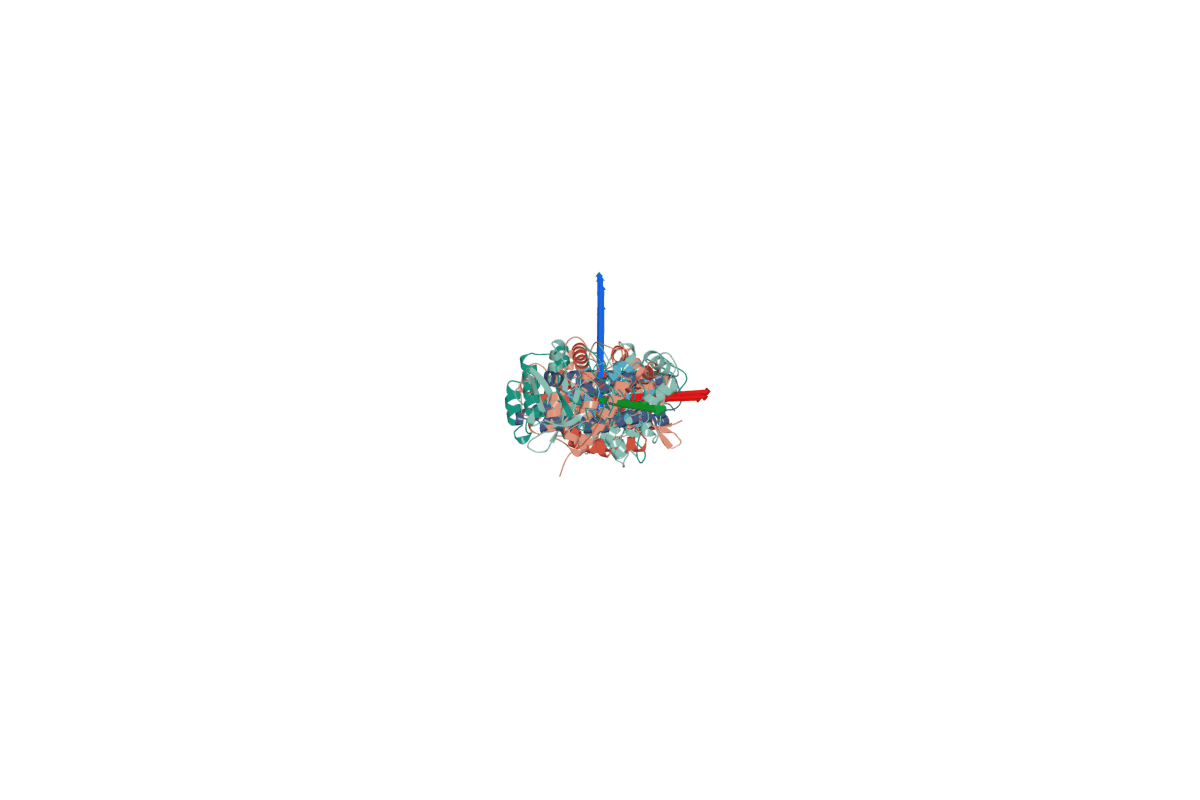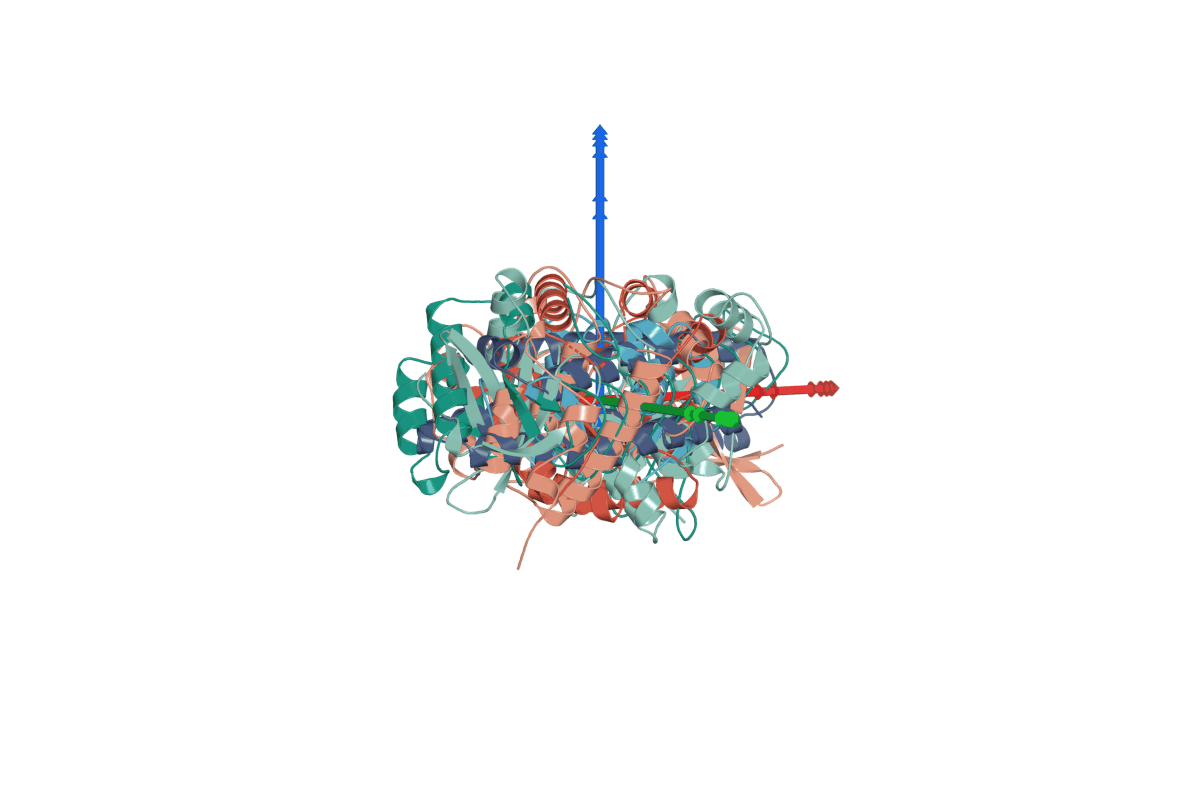
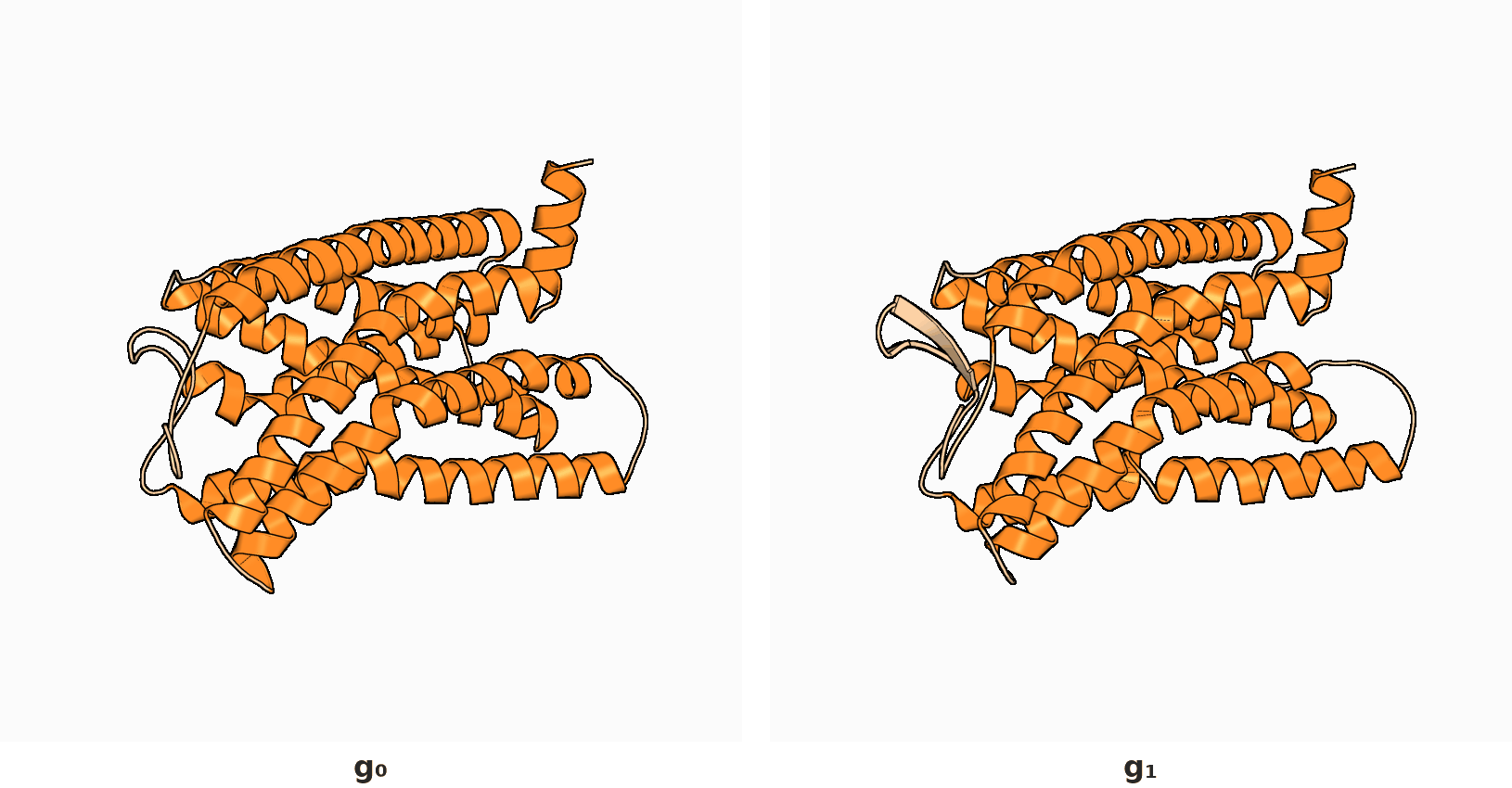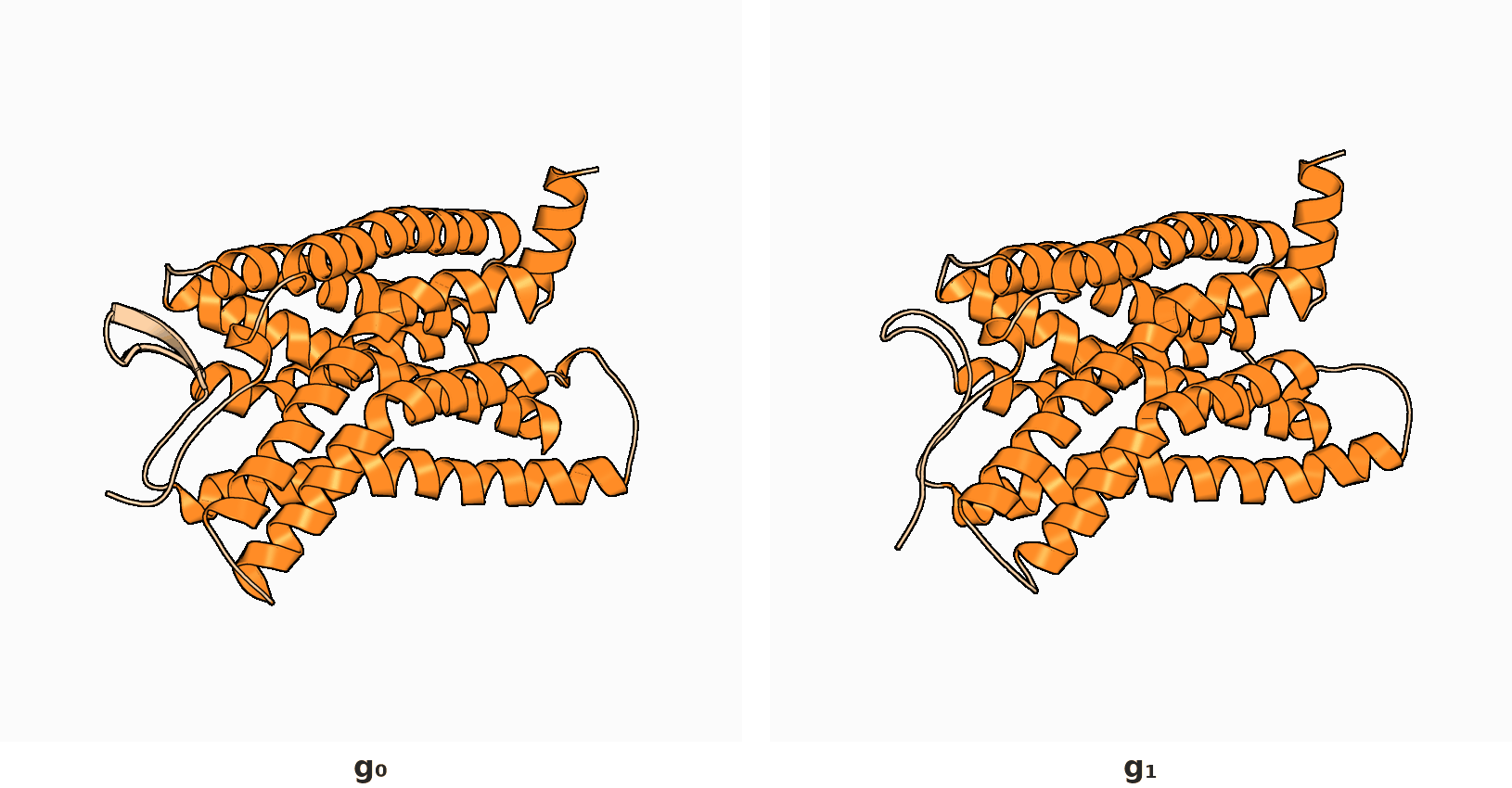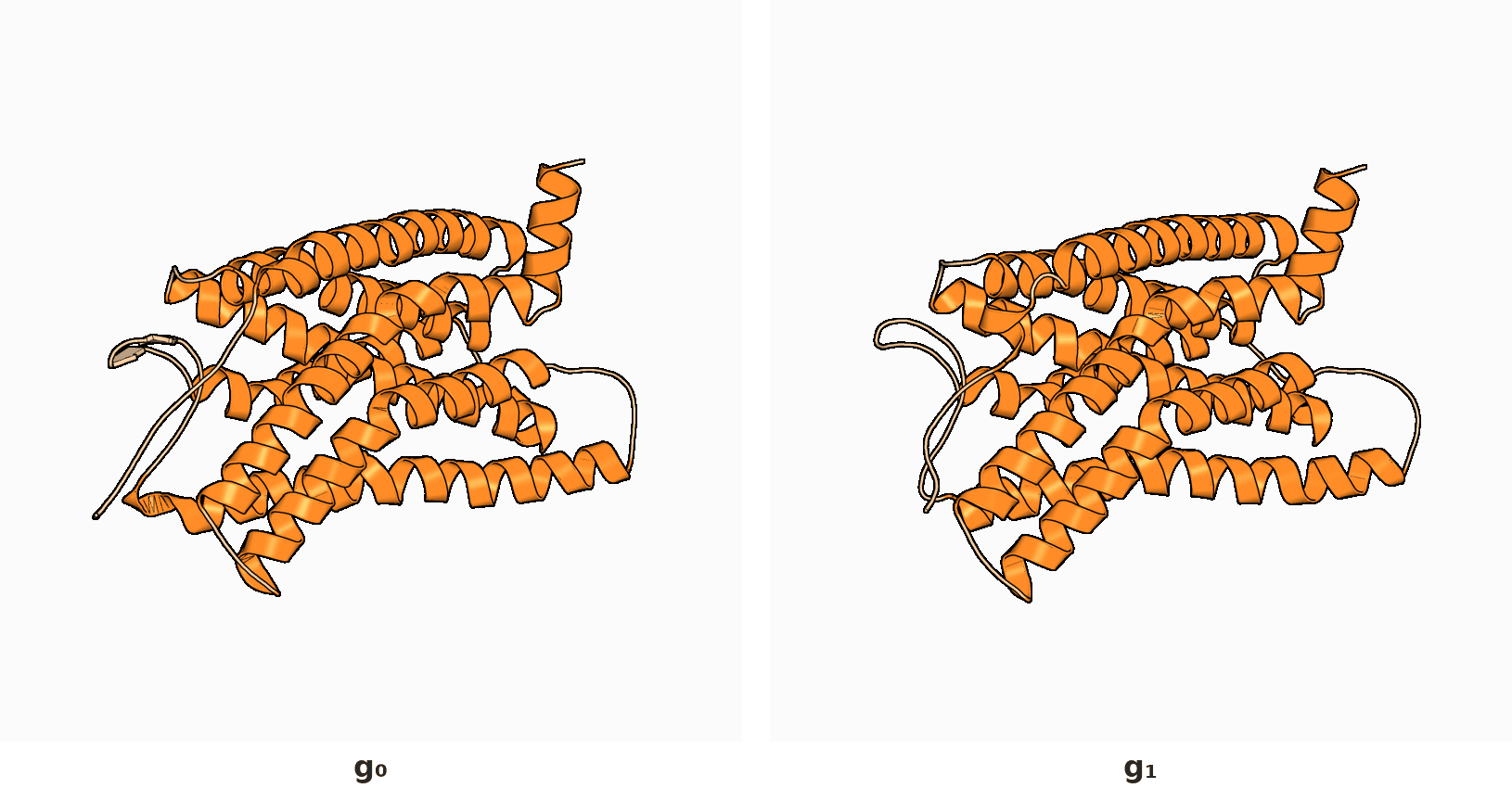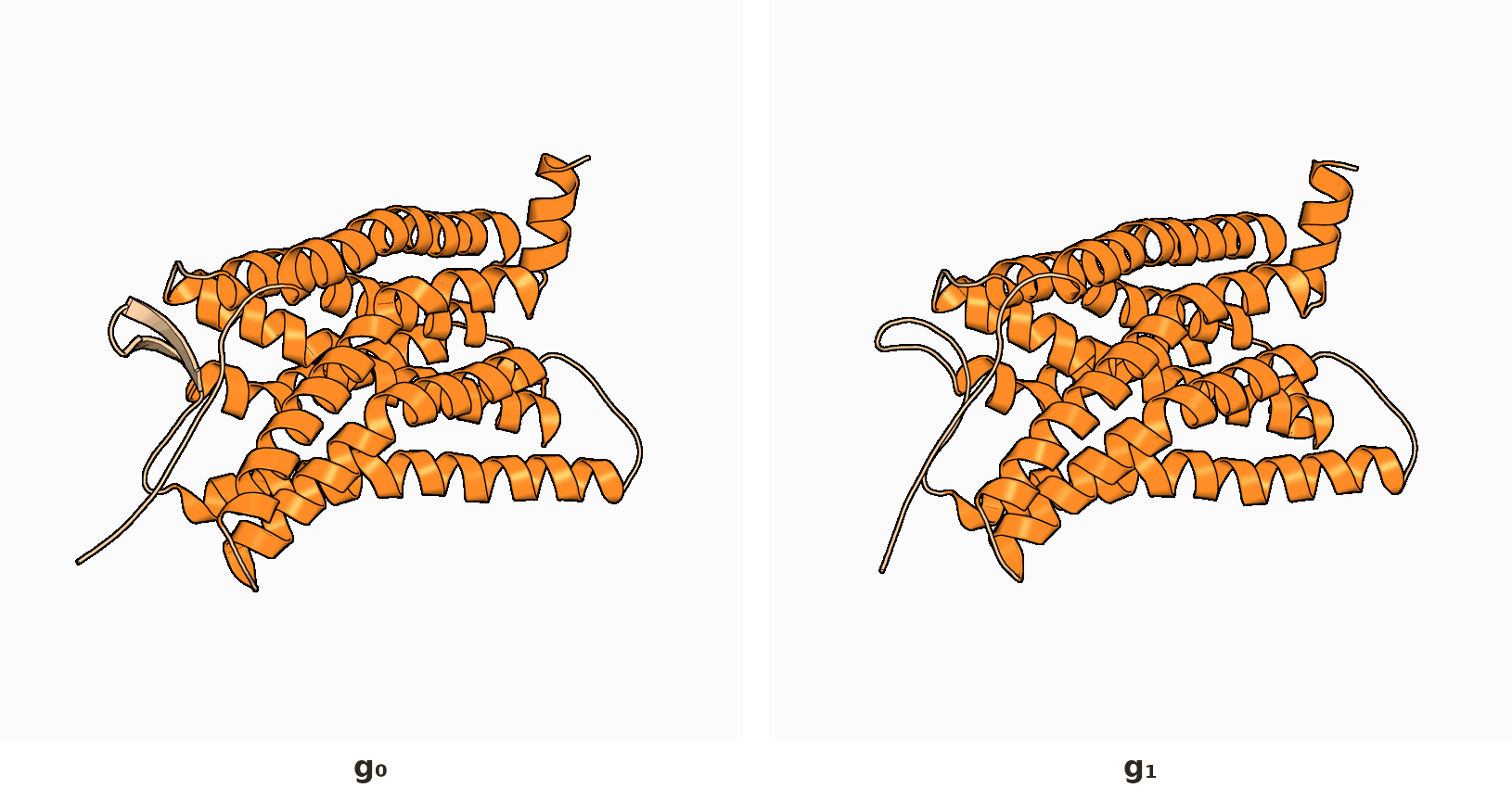
2. RigidSSL learns protein dynamics by maximizing mutual information between two conformations $g_0$ and $g_1$.
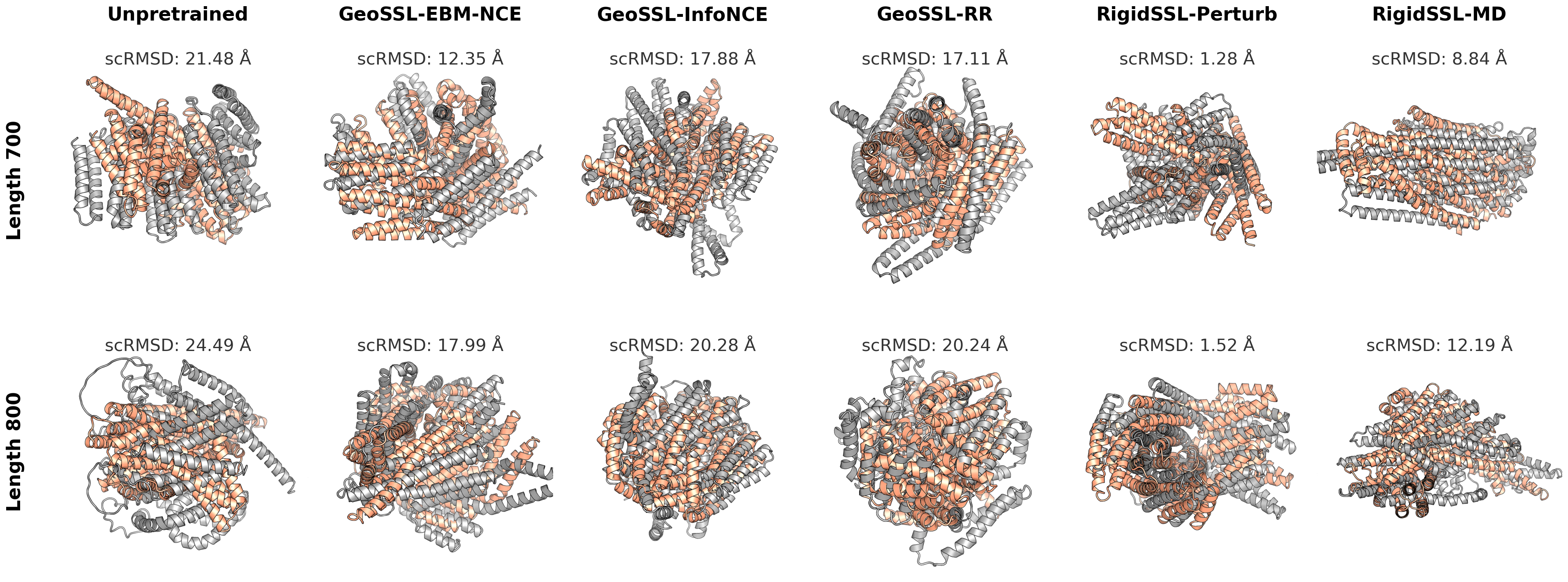
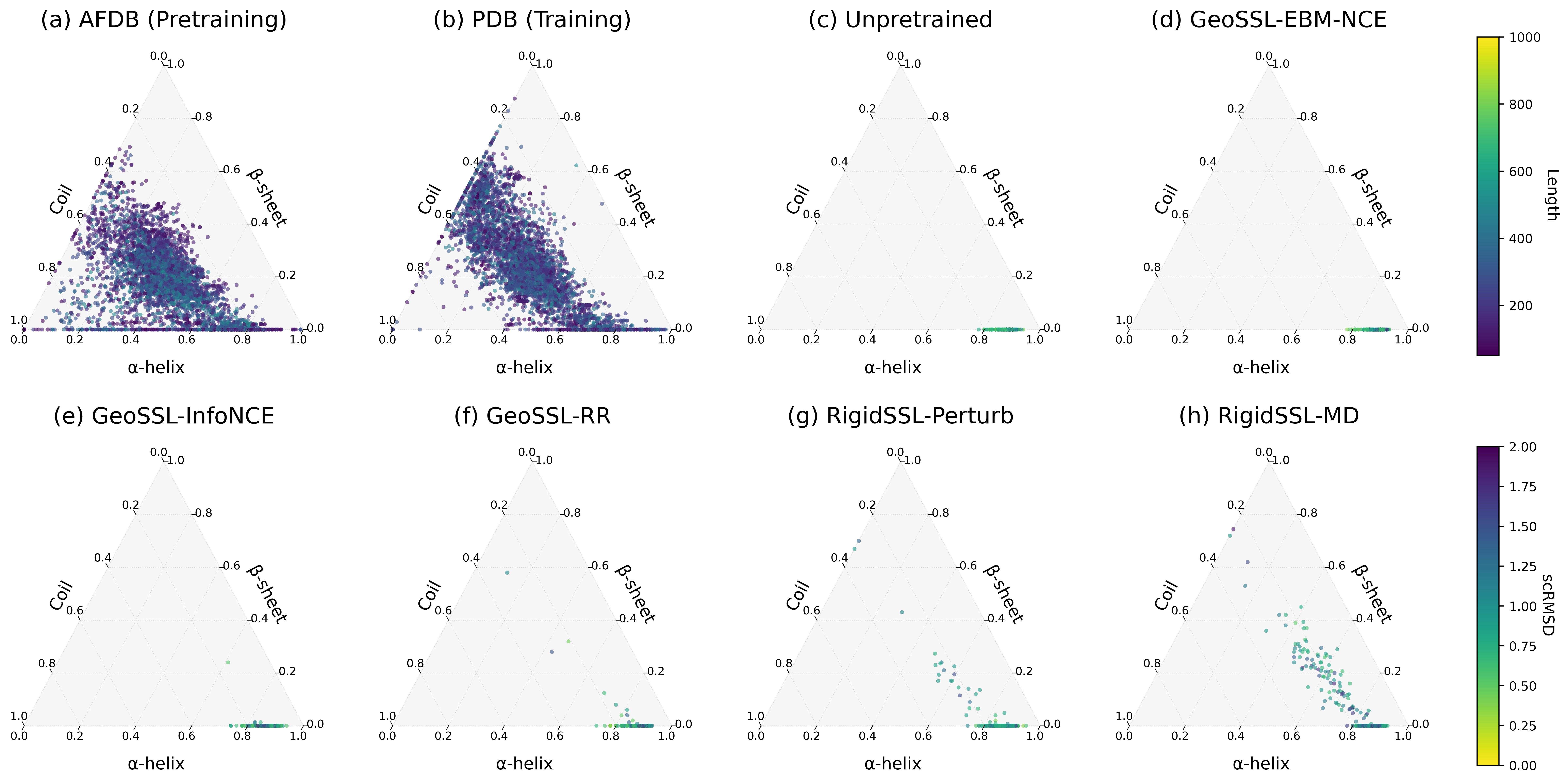
3. RigidSSL variants improve protein structure designability, novelty, diversity, and secondary structure statistics.
4. RigidSSL generalizes to long protein chains of 700-800 residues.